Pedigree drawing,
management, and modelling
Fast pedigree drawing. Twenty published risk-model algorithms. A 1,900-entry catalogue. In the browser.
Fast pedigree drawing. Twenty published risk-model algorithms. A 1,900-entry catalogue. In the browser.

Paper charts get lost in handover. General-purpose diagramming tools don't understand affection fill or proband arrows. Legacy genetics software forces you to re-key family data into a separate BayesMendel calculator — and none of it runs in a browser.
The result: consultation time wasted on tools rather than patients. Incompatible exports between research teams. Families locked out of their own history.
Evagene combines gesture-based pedigree drawing, deep clinical data modelling, Bayesian risk analysis, and universal data exchange — in a single browser application with no installation.
Draw a circle for a female. A square for a male. Connect them. Add diseases with ICD-10 codes. Run BRCAPRO. Export to PDF. That is the entire workflow.
Pick up a stylus — or use your finger, mouse, or trackpad — and draw directly on the canvas. Circles, squares, diamonds, connecting lines. Exactly as you would with pencil and paper.
As you sketch, Evagene recognises each gesture in real time and builds a fully structured genetic model underneath: individuals, relationships, generations, and lineage — ready for annotation, risk analysis, and export.
No menus. No toolbars. No learning curve. If you can draw a pedigree on paper, you can draw one in Evagene.
Drawing a pedigree on iPad with Apple Pencil — gestures become structured data in real time
Our ChatGPT Custom GPT — Evagene Pedigree Builder — turns a natural-language family description into a valid Evagene pedigree JSON file. Describe who is in your family, what conditions they have, and any genetic test results you know about. The GPT asks clarifying questions where family structure is ambiguous and returns a downloadable file.
Load the file into Evagene via File → Load pedigree → Load Evagene file and the pedigree renders in standard notation, ready for risk analysis.
Tell ChatGPT about your family.
Get a JSON pedigree file.
Open it in Evagene.
Requires a paid ChatGPT subscription (Plus / Team / Enterprise) — OpenAI's policy for all Custom GPTs, not an Evagene paywall. ChatGPT helps you produce a valid Evagene file; the analysis runs in Evagene after import.
Quick check — the aunt with ovarian cancer: is she on the maternal side too, and roughly what age at diagnosis?
Here's your pedigree. Three generations on the maternal side with three affected relatives and you as proband. The pattern is suggestive of hereditary breast / ovarian cancer — run BRCAPRO in Evagene for a quantitative read.
Load in Evagene via File → Load pedigree → Load Evagene file.
Illustrative conversation — the GPT asks clarifying questions, then returns a JSON pedigree file.
From gesture-based canvas drawing to implementations of published Bayesian risk-model algorithms and GEDCOM interoperability — a full research and education surface in an interface anyone can learn in minutes.
Freehand-draw circles, squares, and diamonds on the HTML5 canvas. Draw lines to form relationships. Natural as pen and paper, with keyboard accelerators.
Internationally recognised pedigree symbols: affection fill, carrier dots, half-fill, mortality overlays, consanguinity double-lines, proband arrows, and fertility indicators.
Floating draggable panel: identity (8 biological sex options), death status (12 options), lifestyle, contact fields, consent-to-share flags, relationship events — all synced in real time.
Oncology, cardiology, neurology, metabolic, plus 20+ catalogued complex / multifactorial conditions — each entry carries ICD-10, OMIM, inheritance pattern, penetrance tables, heritability, empirical recurrence risks, and carrier frequencies by ancestry. Manage from a dedicated dashboard with presets and custom colours.
BRCAPRO, MMRpro, PancPRO via BayesMendel. Claus, Couch, Frank, Manchester, NICE, Amsterdam II, Bethesda, Gail, and a Tyrer-Cuzick IBIS-style approximation alongside. AD / AR / XLR Mendelian with age-dependent penetrance; X-linked dominant with sex-differential severity; mitochondrial with heteroplasmy scaling; digenic two-locus; imprinting / UPD with parent-of-origin weighting. Polygenic / multifactorial liability-threshold engine covering 20+ complex conditions. One-click CanRisk export.
Ideogram of all chromosomes with genetic markers at genomic positions, colour-coded by test result (positive / VUS / negative), with centromere bands.
Import JSON, GEDCOM 5.5.1, XEG, 23andMe, and images (OCR). Export to JSON, GEDCOM, PNG (up to 4x), SVG, and PDF (A4/A3/Letter/Legal).
Wright's path coefficient calculates consanguinity coefficients automatically. Values display on relationship lines and feed directly into risk models.
Query builder: filter by sex, disease status, age, tests, treatments — across parents, siblings, ancestors, descendants. AND/OR modes. Results highlighted on canvas.
Explore multiple algorithm families against a proband simultaneously. Generate dual-audience educational summaries: one in plain language, one with structured detail. For review, teaching, and documentation — not clinical recommendation.
55+ allergies (severity, IgE, cross-reactivity). 50+ traits (heritability, marker associations) with automatic inference of blood type, Rh factor, and secretor status from 23andMe SNP data.
An educational correlation graph links diseases, traits, clinical-test results, allergies, and markers, surfaced as you annotate — with status-aware test edges (low vs high ferritin). Educational reference, not risk analysis or diagnosis.
Algorithm-based positioning respecting family structure. Undo/redo, copy/paste, lasso select, snap-to-grid, pinch zoom, light/dark theme, configurable fonts.
Generate AI-assisted draft summaries for educational / research review — structural observations, documentation gaps, and discussion prompts, anchored directly to the proband on the canvas. Drafts for human review — not clinical advice, not diagnostic output.
Import 23andMe SNP data and Evagene automatically infers ABO blood type, Rh factor, and secretor status from genotype markers. Traits display as visual cards alongside the pedigree.
Browse, search, and manage your disease collection from a dedicated dashboard. Quick-add presets (heritable cancers, cardiovascular), taxonomy views, custom colours, and ICD-10 filtering.
Auto-generate structured English reports describing the proband and family findings. Rich markdown editor with formatting toolbar, live preview, and anchored canvas notes that follow individuals.
Ready to draw your first pedigree?
Request Alpha AccessResearchers and research cohorts, educators and students, trainees, genealogists, and families documenting their own history. Pick a role below.
Every symbol, fill pattern, and overlay follows standard clinical pedigree conventions — eliminating the ambiguity of hand-drawn charts and the limitations of diagramming tools.
Reproduces peer-reviewed pedigree risk-model algorithms for educational exploration. Demonstrates how each published model behaves under different pedigree inputs. Algorithm implementations for teaching, training, and research — not clinical recommendations.
Reproduces three BayesMendel Bayesian algorithms (BRCAPRO, MMRpro, PancPRO) and nine family-history scoring algorithms on input pedigrees. Demonstrates how each published model behaves under different family-history configurations. Educational exploration only — not clinical outputs.
Reproduces the BayesMendel BRCAPRO algorithm (Parmigiani et al.). Demonstrates how the published Bayesian family-history model behaves under different pedigree inputs.
Reproduces the BayesMendel MMRpro algorithm. Explores how the published Lynch-syndrome model behaves across different pedigree configurations.
Reproduces the BayesMendel PancPRO algorithm. Explores how the published pancreatic-susceptibility model behaves on familial-pancreatic-cancer pedigree inputs.
Each card below reproduces a specific peer-reviewed algorithm on pedigree input for educational exploration. Evagene does not state what the output means clinically — that is a matter for the published literature and qualified professionals.
Reproduces the CASH-derived algorithm for educational exploration of how a family-history-only Bayesian model behaves under varied pedigree inputs.
Reproduces the published logistic-regression algorithm. Demonstrates how the model behaves under different age, ovarian, and Ashkenazi inputs.
Reproduces the empirical scenario-based frequency tables published from the Myriad dataset.
Reproduces the published Manchester point-based scoring algorithm. Algorithm demonstration; interpretation of output remains with the published literature.
Reproduces the categorisation logic published in NICE CG164 and NG101 for educational reference. The guideline itself, not this software, is the source of any action.
Reproduces the five-point Amsterdam II evaluation logic. Flags the pedigree-derivable criteria for teaching purposes.
Reproduces the revised Bethesda criteria evaluation logic on input pedigrees. Algorithm reproduction for teaching reference.
Reproduces the NCI BCRAT computation for educational exploration of how reproductive and family-history inputs drive the published algorithm.
Reproduces the published Tyrer-Cuzick 2004 algorithm. Approximation, not the official IBIS binary (full IBIS coefficients are not publicly released).
Implementations of seven peer-reviewed pedigree inheritance models for educational exploration. Demonstrates how each behaves on pedigree input — autosomal dominant, autosomal recessive, X-linked recessive, X-linked dominant with sex-differential severity, mitochondrial maternal transmission, digenic two-locus interaction, and imprinting / UPD with parent-of-origin weighting.
mtDNA mutations transmitted through the maternal line. Sex-differential penetrance (e.g. LHON male preponderance). Phenotype severity scales with mutant-load (heteroplasmy).
Phenotype requires a simultaneous variant at two separate loci. An affected (Aa;Bb) × unaffected (aa;bb) partnership yields 25% affected offspring — the canonical digenic ratio. Supports both_het, one_het_one_hom, both_hom configurations.
Four molecular mechanisms modelled (deletion, UPD, imprinting-centre defect, point mutation). Recurrence risk is weighted by mechanism; for IC defects, the parent-of-origin rule applies (transmitting parent determines which syndrome phenotype).
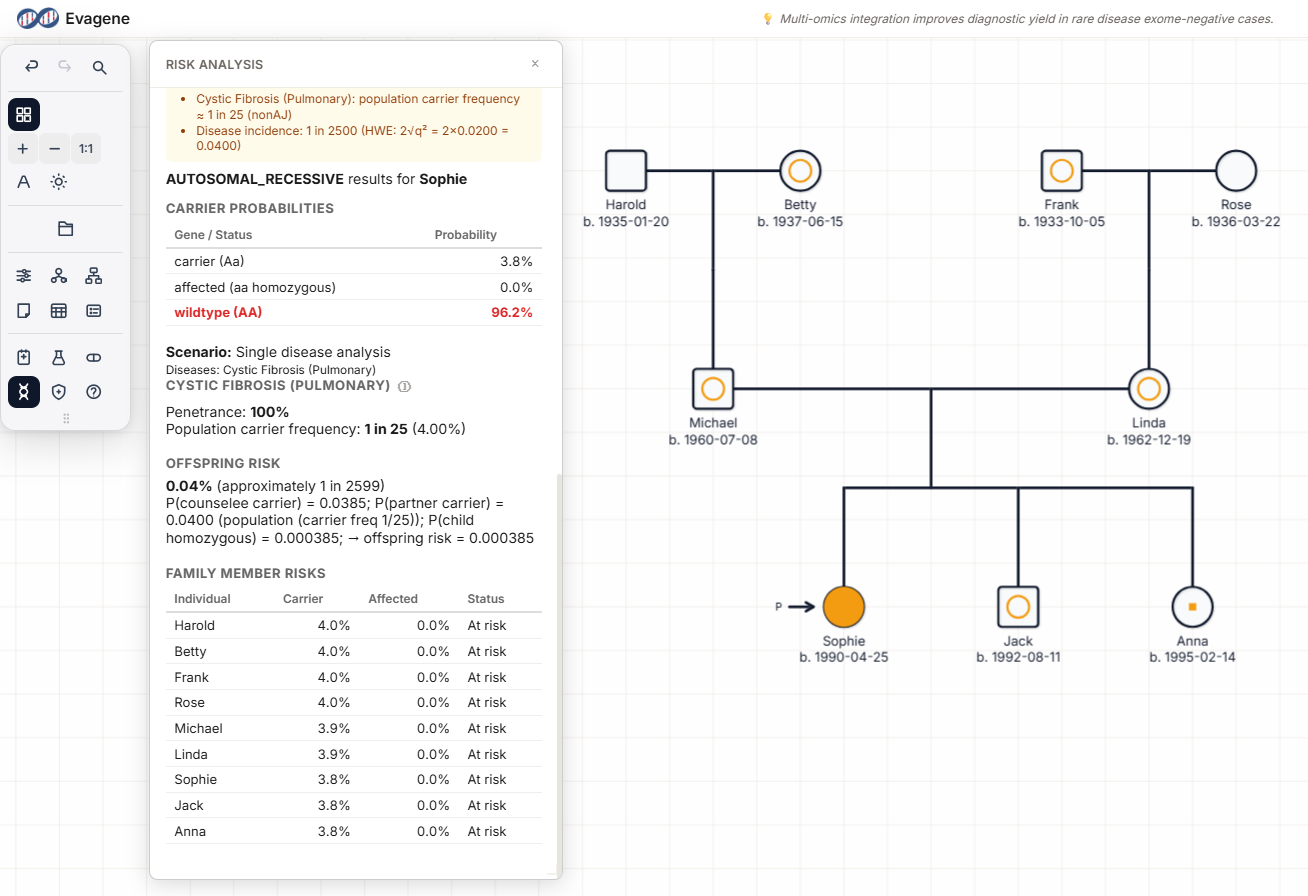
Hardy-Weinberg prior-carrier algorithms are notoriously easy to misuse by outputting a "general population" number for someone whose ancestry is not recorded. Evagene's implementation refuses that. If an individual has no ancestry recorded, the algorithm returns no personalised number for that individual. When ancestry is recorded, outputs are attributed to one of three explicit algorithm paths:
Direct published carrier frequency for that ancestry. Cystic fibrosis in Northern European, ~1 in 25. Tay-Sachs in Ashkenazi Jewish, ~1 in 30.
Proportion-weighted combination across relevant populations when ancestry is mixed. Per-population contributions stay visible in the row.
Derived as 2pq where no population-specific carrier frequency is published. Flagged as an estimate, not a published figure.
With no ancestry recorded, the algorithm returns no value for that individual. By design the software never outputs a single "general population" figure as though it applied to an identifiable person. The output is a property of the algorithm — not an opinion from the software about any individual.
Reproduces the published liability-threshold algorithm (Carter 1961, Falconer 1965, Reich / James / Morton 1972) on pedigree input, with the empirical family-study tables (Smith, Carter, Harper) where published. Demonstrates how the algorithm behaves across common complex disorders that cluster in families but do not follow Mendelian patterns. Algorithm reproduction for teaching and research — not clinical output.
Evagene does not compute BOADICEA. It writes a ##CanRisk 2.0 pedigree file (BOADICEA v4 column set) that the user downloads and uploads at canrisk.org, where the computation happens off-platform under University of Cambridge licence.
BOADICEA is licensed by the University of Cambridge; third-party web-service integration requires a separate commercial licence. Evagene therefore does not bundle BOADICEA — the file-export approach puts all computation and responsibility off-platform, where it belongs.
Import from clinical systems, genealogy platforms, and consumer genomics. Export publication-ready output in any format.
| Format | Import | Export | Notes |
|---|---|---|---|
| JSON | ✓ | ✓ | Full snapshot; replace or add mode |
| GEDCOM 5.5.1 | ✓ | ✓ | Ancestry, FamilySearch, Gramps compatible |
| XEG | ✓ | — | Legacy Evagene v1 XML migration |
| 23andMe | ✓ | — | Genotype, ancestry, traits, health history, conflict resolution |
| Image | ✓ | — | OCR + shape detection from chart photos |
| ChatGPT Custom GPT | ✓ | — | Describe your family to the Evagene Pedigree Builder — get back importable JSON (paid ChatGPT tier required) |
| PNG | — | ✓ | 1x–4x scale, optional transparency |
| SVG | — | ✓ | Vector for publication-quality output |
| — | ✓ | A4/A3/Letter/Legal, portrait/landscape/auto | |
| CanRisk / BOADICEA v4 | — | ✓ | ##CanRisk 2.0 header; tab-separated, BOADICEA v4 columns; upload at canrisk.org (BOADICEA not bundled — licensed by Univ. of Cambridge) |
Build integrations, connect AI agents, push data to external systems, and embed pedigree diagrams in your own applications. Evagene's platform layer turns pedigree intelligence into infrastructure.
Long-lived, scoped, rate-limited keys for programmatic access to the full Evagene API. SHA-256 hashed at rest. Configurable per-minute and per-day limits.
Supply your own Anthropic or OpenAI API key for AI analysis. Encrypted at rest with Fernet. No quota limits. Choose your provider and model.
Push HMAC-SHA256 signed notifications when pedigrees change. Connect to EHR systems, LIMS, Slack, or any endpoint that accepts HTTP POST.
Reusable custom AI prompt templates with variable injection. Design analyses for pharmacogenomics, reproductive risk, research data extraction, or patient-friendly summaries.
Let AI agents work with pedigrees via the Model Context Protocol. 15 tools for Claude Desktop, custom agents, or any MCP-compatible client. Direct database access, no HTTP overhead.
Drop-in pedigree diagrams for patient portals, research dashboards, or any website. Three embedding modes: iframe, raw SVG, or a JavaScript snippet.
Evagene has been cited in clinical and genetic studies published in the Journal of Clinical Laboratory Analysis, Bioengineering, and Health Science Reports, covering Marfan syndrome, nephronophthisis, retinitis pigmentosa, and molecular diagnostics.
Every disease, trait, allergy, clinical test, genetic marker / gene, and treatment in Evagene has a dedicated help page with catalogue metadata, cross-references to related entries, and authoritative external links — LOINC for tests, NCBI Gene / OMIM / ClinVar for markers, RxNorm / BNF / DrugBank for treatments.
Covers the central dogma (DNA → RNA → protein), chromosome structure, Mendelian inheritance patterns, and how mutations lead to disease. Written for readers with no prior genetics background. Includes diagrams of DNA replication, meiosis, and autosomal vs. X-linked inheritance.
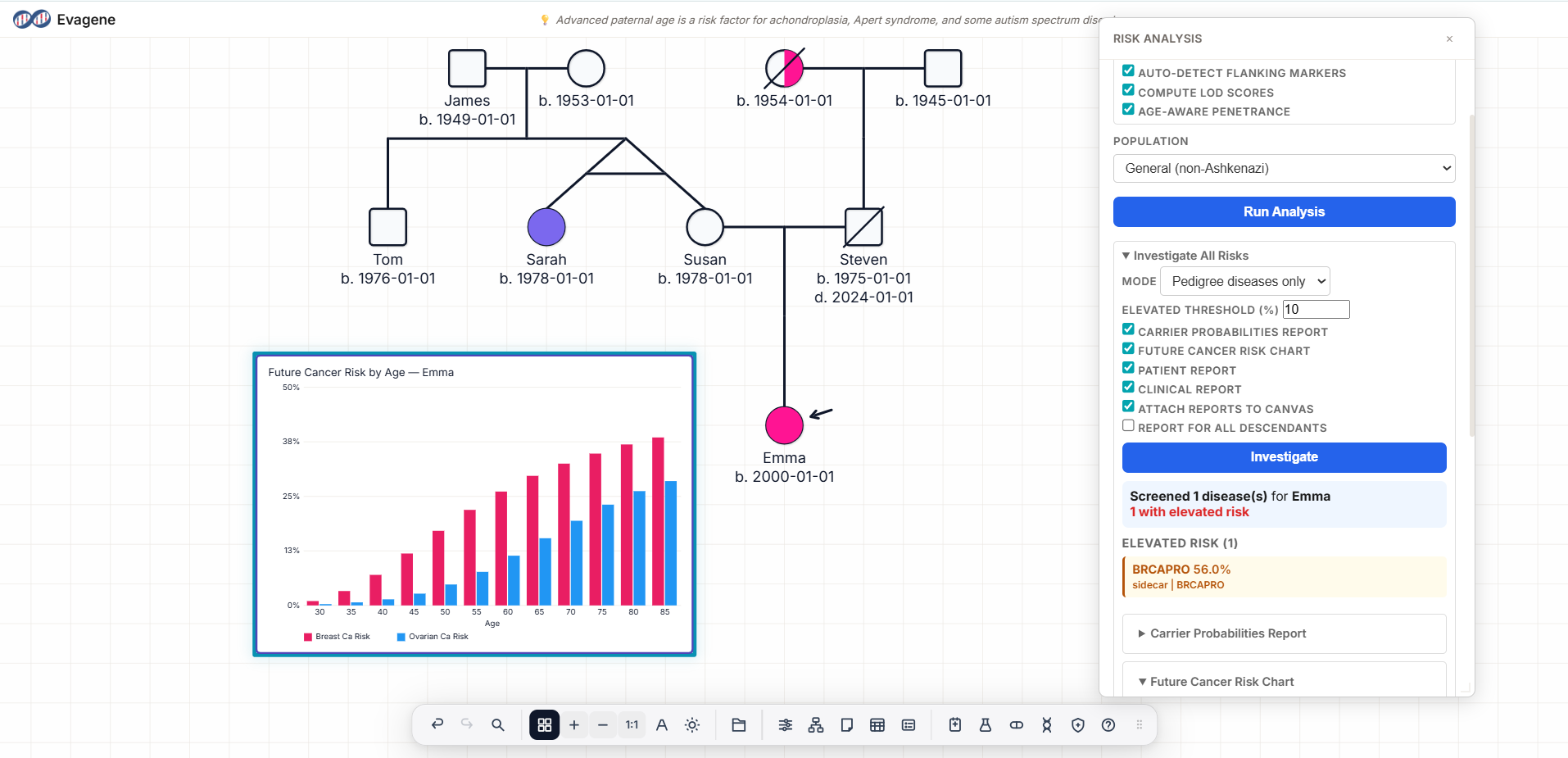
BRCAPRO uses Bayesian updating with the BayesMendel R package to estimate BRCA1/BRCA2 carrier probability from family pedigree data. Supports General, Ashkenazi Jewish, and Italian population frequencies. Published literature discusses ≥10% and ≥25% as notable thresholds; ≥25% is strongly suggestive of hereditary breast and ovarian cancer syndrome. These are educational references — not clinical recommendations from this software.
ICD-10: C50. OMIM: 114480. Inheritance: multifactorial with high-penetrance genes (BRCA1, BRCA2, TP53, PALB2). Lifetime risk ~12% general population; up to 72% for BRCA1 carriers. Evagene tracks laterality, ER/PR/HER2 status, grade, stage, and age at diagnosis per individual.
M/F/U to add individuals, P for partner, D for disease palette, N for notes, G for genetics panel. Ctrl+Z/Y for undo/redo, Ctrl+C/V for copy/paste, Ctrl+A to select all, F2 to edit name, Delete to remove. Space+drag to pan, scroll to zoom, Ctrl+G for snap-to-grid.
What Evagene is. A browser-based platform for drawing, managing, and exploring family pedigrees. Used by clinicians, counsellors, researchers, educators, students, and families for teaching, training, research, and structured family-history documentation.
What Evagene is not. Evagene is not a medical device. It is not intended to diagnose, prevent, monitor, predict, treat, or manage disease; determine eligibility for screening, testing, referral, or treatment; or replace professional clinical judgement. Outputs are illustrative and for educational and research purposes only.
Two risk-model caveats that apply everywhere on this site: Tyrer-Cuzick output is an IBIS-style approximation of the published Tyrer / Duffy / Cuzick 2004 algorithm, not the official IBIS Breast Cancer Risk Evaluator binary. BOADICEA is not bundled — licensed by the University of Cambridge; Evagene exports a ##CanRisk 2.0 pedigree file that the user uploads at canrisk.org.
Be among the first to use Evagene. Alpha members receive early access, direct input on the roadmap, and priority support from the development team.
We'll notify you as soon as alpha access opens. In the meantime, share Evagene with colleagues who work with pedigrees.
Join clinicians, researchers, and families already on the waiting list. Early members shape the roadmap and receive priority support.
Request Alpha Access